CONTENTS
2017 Fall
RNA web demo
- 12/05/2017 - Task Initiallization
- 12/06/2017 - Getting Familiar
- 12/07/2017 - Layout
- 12/08/2017 - Arc draw in canvas
- 12/10/2017 - Rearrange Data by Seq
- 12/11/2017 - Learn JavaScript to draw
- 12/12/2017 - jQuery night
- 12/13/2017 - File Reading Problem
- 12/14/2017 - Staged Success
- 12/26/2017 - Fully Functioned
- 12/28/2017 - P/R/F and other optimizations
- 12/29/2017 - Plot in html
- 12/30/2017 - Fulfill many tough requests
- 12/31/2017 - My own updates in format
- 01/01/2018 - Problem shooting and ssh-key research
- 01/02/2018 - Trial of less file loading & Enable re-draw by chart click
- 01/17/2018 - Interactive web demo with flask
- 01/18/2018 - File upload to server & stylish page
Pseudoknot
12/05/2017 Tue
Meeting with Prof. Huang and Dezhong, on the project of RNA sequence pairing.
RNA paring for pseudoknot-free mode is perfectly improved in linear solution. Next step is to optimize different types of crossing, from $O(n^3)$ to $O(n^6)$, to linear.
The group also needs a web demo for visual illustration, especially for beam tunning. So I took the task for web development. Everything needs to study from the very beginning.
Back to subcontents RNA web demo
12/06/2017 Wed
Check RNA_visual at Dezhong’s file on ironcreek: ssh liukaib@ironcreek.eecs.oregonstate.edu
. Then copy them to my local:
scp -r liukaib@ironcreek.eecs.oregonstate.edu:/scratch/dengde/RNA_visual /Users/zhangyilin/0_pipi/LKB_OSU/Research/RNA
Borrow the source code from Vienna RNAfold WebServer. I need to modify many contents then deploy html file to my own webpage on engr:
# at OSU
scp /Users/zhangyilin/0_pipi/LKB_OSU/Research/RNA/web_demo/RNA_pair_web.html liukaib@flop.engr.oregonstate.edu:/nfs/stak/users/liukaib/public_html
# out of OSU
scp -r /Users/zhangyilin/0_pipi/LKB_OSU/Research/RNA/web_demo/rearranged_results/ liukaib@flop.engr.oregonstate.edu:/nfs/stak/users/liukaib/public_html
Back to subcontents RNA web demo
12/07/2017 Thu
Layout:
graph LR
contrafold-->contrafold_linear
vienna-->vienna_linear
Back to subcontents RNA web demo
12/08/2017 Fri
- Use
canvasin html5 to draw arcs for the pairing demostration.//get canvas object var canvas = document.getElementById("myCanvas"); //check if current explorer support Canvas object, to avoid sytax error in some html5-unfriendly explorers. if(canvas.getContext) { //get corresponding CanvasRenderingContext2D object(pen) var ctx = canvas.getContext("2d"); ctx.beginPath(); //starting drawing a path ctx.moveTo(p1.x, p1.y); //assign the start position of drawing //draw an arc of radius r, tangent with 2 sides consisted by p1 to (x0,y0) and (x0,y0) to p2. ctx.arcTo(x0, y0, p2.x, p2.y, r); //or, draw a line from p1 to p2 ctx.lineTo(p2.x,p2.y); ctx.strokeStyle = color; //set arc to a color assigned by a string color ctx.stroke(); }; - Here is the example:
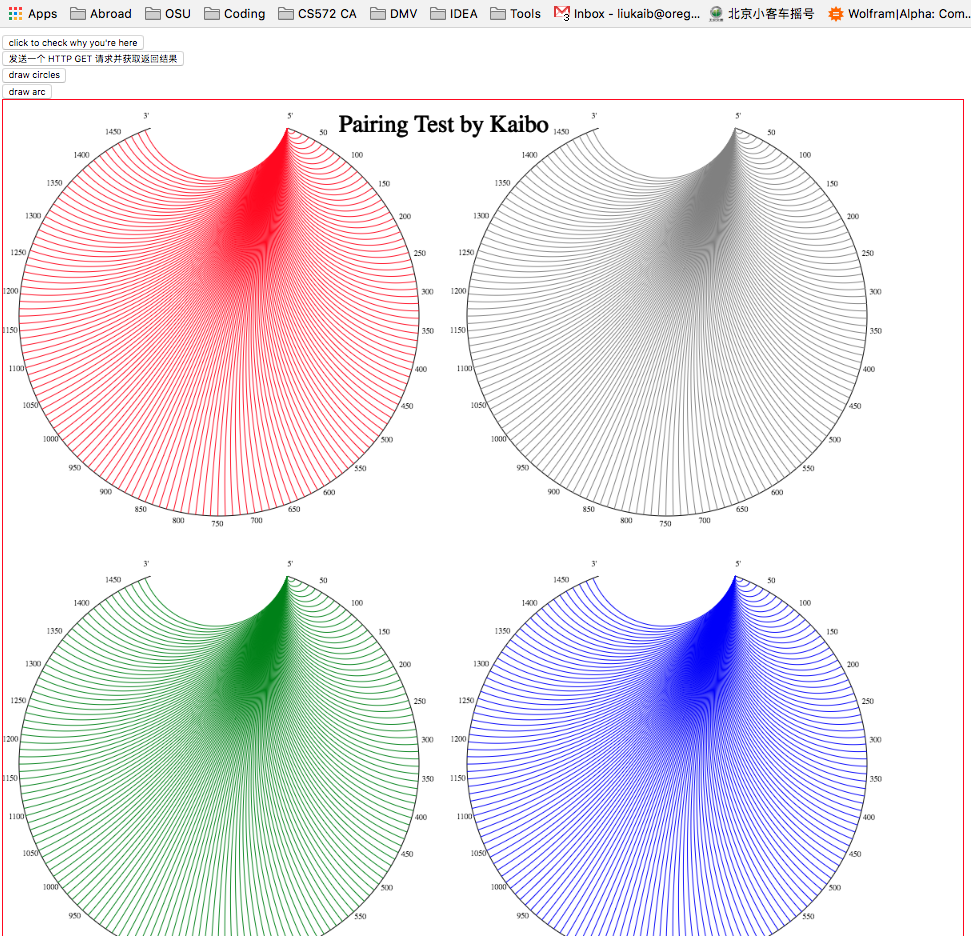
Back to subcontents RNA web demo
12/10/2017 Sun
In Dezhong’s README, data are listed like this:
Original data (sequences):
Mathewsdata.16s.seq
Mathewsdata.23s.seq
Gold:
Mathewsdata.16s.ref
Mathewsdata.23s.ref
Contrafold result:
Mathewsdata.16s.contrafoldres
Mathewsdata.23s.contrafoldres
Vienna result:
Mathewsdata.16s.viennares
Mathewsdata.23s.viennares
Linear Parser result using Contrafold model:
linearcontrafold/run_16s/*
linearcontrafold/run_23s/*
Linear Parser result using Vienna model:
linearvienna/run_16s/*
linearvienna/run_23s/*
Finishing rearrange_by_seq.py, rearrangement of pairing, put all the results of a single sequence (#4 of 16s, for example) into a data file, named combine_16s.seq04.
There are always 1656 lines(seq*2 + ref*2 + cf*2 + vn*2 + linearcf*206*4 + lineavnf*206*4) in each data file if category of beams is 206 (1,2,..200,300,..,800).
The structure of the file is:
seq * 2 lines -----> L1
ref * 2 lines -----> L3
cf * 2 lines -----> L5
vn * 2 lines -----> L7
linearcf.beam001 * 4 lines -----> L11 (with 2 lines of information above)
lineavnf.beam001 * 4 lines -----> L16 (with 2 lines of information above)
linearcf.beam002 * 4 lines
lineavnf.beam002 * 4 lines
...
linearcf.beam00i * 4 lines -----> L(8*i+3) (with 2 lines of information above), if i > 200, L[(i/100+198)*8+3]
lineavnf.beam00i * 4 lines -----> L(8*i+7) (with 2 lines of information above), if i > 200, L[(i/100+198)*8+7]
通宵
Back to subcontents RNA web demo
12/11/2017 Mon
Think about methods to make javascript to read data files like combine_16s.seq04 to draw graph.
半通宵
Back to subcontents RNA web demo
12/12/2017 Tue
jQuery.get( url [, data ] [, success(data, textStatus, jqXHR) ] [, dataType ] )
描述: 使用一个HTTP GET请求从服务器加载数据。
It’s an abbreviation of ajax, equalent to:
$.ajax({
url: url,
data: data,
success: success,
dataType: dataType
});
Usage:
jQuery.get('http://localhost/foo.txt', function(data) {
var myvar = data;
});
ps: jQuery.get()=$.get()
HTML:
element: a pair of tags and all those between it
- has closing tag:
<h1>,<button>,<a>,<div>,<body>,<html> - closing tag is optional:
<p>,<option>; - no closing tag:
<br>,<img>
attribute: in the openning tag(class, id, style, title, lang)
class 属性可以多用 class=” “ (引号里面可以填入多个class属性)
id 属性只能单独设置 id=” “(只能填写一个,多个无效)
jQuery selector:
element selector $("p")
id selector $(#id)
class selector $(.class)
Back to subcontents RNA web demo
12/13/2017 Wed
Solved an interested problem:
When I use $.get(), the URL should be an open access domain, otherwise there will be an error No 'Access-Control-Allow-Origin' header is present on the requested resource. Origin 'null' is therefore not allowed access..
The browser usually allows a request in the same origin for security reasons. CORS is designed for cross-domain request. Or simply without CORS:
- If I want to
$.get()a file on server, the html file should also be run on the server. - If I want to
$.get()a file on github, I can run html both locally or on server.
$(document).ready(function(){
// jQuery...
});
is equivalent to
$(function(){
// jQuery...
});
Back to subcontents RNA web demo
12/14/2017 Thu
Use json.dump(a:b) in python and $.getJSON(URL,function(data){...}) in js, to make file data reable for html.
But I cannot pass variables out in this getJSON.
This happens because that callback function (function(data) {…}) runs later when the response comes back…because it’s an asynchronous function. Instead use the value once you have it set, like this:
…
After finishing all the configuration, I can draw 4 beautiful graphs for a default data file(a specific sequence) on the webpage.
Transfer html, js, arc_pairing_json.py, rearrange_by_seq.py, and data file pairing_for_js to server:
scp /Users/zhangyilin/0_pipi/LKB_OSU/Research/RNA/web_demo/arc_pairing_json.py liukaib@flop.engr.oregonstate.edu:/nfs/stak/users/liukaib/public_html/
scp /Users/zhangyilin/0_pipi/LKB_OSU/Research/RNA/web_demo/demo_myScript.js liukaib@flop.engr.oregonstate.edu:/nfs/stak/users/liukaib/public_html/
scp /Users/zhangyilin/0_pipi/LKB_OSU/Research/RNA/web_demo/demo_json+canvas.html liukaib@flop.engr.oregonstate.edu:/nfs/stak/users/liukaib/public_html/
scp -r /Users/zhangyilin/0_pipi/LKB_OSU/Research/RNA/web_demo/pairing_for_js/ liukaib@flop.engr.oregonstate.edu:/nfs/stak/users/liukaib/public_html/
scp /Users/zhangyilin/0_pipi/LKB_OSU/Research/RNA/web_demo/rearrange_by_seq.py liukaib@flop.engr.oregonstate.edu:/nfs/stak/users/liukaib/public_html/
My demo is available on http://web.engr.oregonstate.edu/~liukaib/demo_json+canvas.html. I sent an email to my boss for my update.
凌晨7点睡。
Back to subcontents RNA web demo
12/26/2017 Tue
Start working at midnight.
Add the functionality of slide bar value call back, then extract seq series and seq number from seletion.
Learn the usage of <span>, interesting.
The final demo can be found at my OSU homepage.
Finish the work at 5:50am.
Back to subcontents RNA web demo
12/28/2017 Thu
As boss asked, I need to:
- Change the max value on slide bar(from 206 to 800) and triple the length of the bar.
- I add hash marks and labels on the bar, however, currently, no browser fully supports these features. Firefox doesn’t support hash marks and labels at all, for example, while Chrome supports the hash marks but doesn’t support labels. Source
- Add sequence names to corresponding dropdown manu items.
- Dezhong updated an
Mathews.clean.txton his RNA_visual directory. There are 3867 sequences(11571 lines) in the file. I need to match all the 27 (22 in 16s and 5 in 23s) by searching sequence string. Results are attached later.
- Dezhong updated an
- Add PPV (precision), Sensitivity (recall), F-score, and number of (predicted) pairs for each plot under subtiles.
$P=\frac{\#\ of\ correctly\ predicted pairs}{\#\ of\ predicted\ pairs}$$R=\frac{\#\ of\ correctly\ predicted\ pairs}{\#\ of\ gold\ pairs}$$F=\frac{2PR}{P+R}$- At first I calculate these 3 parameters from the number of my 3 tpye of pairs(
wrong,hit,missing). Actually the number$hit+missing\neq \#\ of\ ref\ pairs$, because there is a slip in pairing(useagreeas equivalence).- For example,
(1,30)in result and(1,29),(2,30)in ref, finally(1,30)will be counted as blue(hit) while no grey pair. - So I need to pass more para into json format data file for js, detailed in JSON file layout.
- For example,
- Update
agreefunction from Dezhong’s fix in myarc_pairing.pyfor correct blue pair statistics, after 1.5h voice call. We both had a bug in counting correct-predicted pairs. Now we get the same corresponsive P/R/F result with Dezhong. - Update algorithm names and performance names, I need to fix text position in canvas, which is time consuming:
- P/R/F -> PPV/Sensitivity (F)
- CONTRAfold -> CONTRAfold MFE
- Vienna -> Vienna RNAfold
- Linear CONTRAfold -> LinearFold-C
- Linear Vienna -> LinearFold-V
- Future task: scalable vector graph(SVG)
domain page
I figured out that if I want to load a default .html file on my domain instead of using an url ending with .html, I just need to rename the page file to index.html.
Sequence names
16s-seq01: 1490 archiveII/16s_Z.mays.ct
16s-seq02: 1552 archiveII/16s_B.subtilis.ct
16s-seq03: 1542 archiveII/16s_E.coli.ct
16s-seq04: 1474 archiveII/16s_H.volcanii.ct
16s-seq05: 1995 archiveII/16s_D.melanogaster.ct
16s-seq06: 1793 archiveII/16s_T.gondii.ct
16s-seq07: 954 archiveII/16s_H.sapiens.mito.ct
16s-seq08: 1537 archiveII/16s_B.burgdorferi.ct
16s-seq09: 1564 archiveII/16s_A.pyrophilus.ct
16s-seq10: 1525 archiveII/16s_P.staleyi.ct
16s-seq11: 1552 archiveII/16s_C.psittaci.ct
16s-seq12: 950 archiveII/16s_C.elegans.ct
16s-seq13: 1804 archiveII/16s_F.ananassa.ct
16s-seq14: 1562 archiveII/16s_T.maritima.ct
16s-seq15: 1474 archiveII/16s_C.reinhardtii.chloro.ct
16s-seq16: 1974 archiveII/16s_M.polymorpha.ct
16s-seq17: 1492 archiveII/16s_A.fulgidus.ct
16s-seq18: 1870 archiveII/16s_H.sapiens.ct
16s-seq19: 1800 archiveII/16s_S.cerevisiae.ct
16s-seq20: 1497 archiveII/16s_P.occultum.ct
16s-seq21: 1200 archiveII/16s_C.reinhardtii.mito.ct
16s-seq22: 1452 archiveII/16s_G.intestinalis.ct
23s-seq01: 2968 archiveII/23s_H.pylori.ct
23s-seq02: 2915 archiveII/23s_T.thermophilus.ct
23s-seq03: 2927 archiveII/23s_B.subtilis.ct
23s-seq04: 2923 archiveII/23s_S.aureus.ct
23s-seq05: 2904 archiveII/23s_E.coli.ct
JSON file for js
combine_pairing_16s.seq04
{"result":[N,seq, ----->index: 0-1
[P,R,F],cf_missing, cf_hit, cf_wrong, ----->index: 2-5
[P,R,F],vn_missing, vn_hit, vn_wrong, ----->index: 6-9
[P,R,F],linearcf_b01_missing, linearcf_b01_hit, linearcf_b01_wrong, ----->index: 10-13
[P,R,F],linearvn_b01_missing, linearvn_b01_hit, linearvn_b01_wrong, ----->index: 14-17
[P,R,F],linearcf_b02_missing, linearcf_b02_hit, linearcf_b02_wrong, ----->index: 18-21 (8*i+2~8*i+5,i is the beam number)
[P,R,F],linearvn_b02_missing, linearvn_b02_hit, linearvn_b02_wrong, ----->index: 22-25 (8*i+6~8*i+9,i is the beam number)
...
[P,R,F],linearcf_b800_missing, linearcf_b800_hit, linearcf_b800_wrong, ----->index: 1238-1241 [(i/100+198)*6+2~(i/100+198)*6+5,i is the beam number which is > 200]
[P,R,F],linearvn_b800_missing, linearvn_b800_hit, linearvn_b800_wrong, ----->index: 1242-1245 [(i/100+198)*6+6~(i/100+198)*6+9,i is the beam number which is > 200]
]}
The new pairing data files will listed by seq number(16s * 22, and 23s *5), 27 files in total.
Structure of js drawing
$.getJSON(seqFile, function(data,status) {
drawFrame();
drawCircle(); // open circle;
drawCircleMarks();// scale adjustable in para 'range'
fillCircles();
fillCircle();
drawArc() in for loop;
}
Back to subcontents RNA web demo
12/29/2017 Fri
New Requst:
“one more easy thing you could do on your current version is:
- add one plot (for both contrafold row and vienna row) where the y axis shows P, R, F and the x axis is the beam size.
- add another plot (for both contrafold row and vienna row) where the x axis is P and y axis is R.
These two tasks can be described as P/R/F-beam plot and R-P plot. I have no clue about ploting in HTML. Research shows avaiable methods are below:
- :sleeping: CanvasJS.Chart, ‘trial’ watermark in free version
<script src="https://canvasjs.com/assets/script/canvasjs.min.js"></script> - :ok_hand: google.charts, powerful and open source
<script type="text/javascript" src="https://www.gstatic.com/charts/loader.js"></script>
Ways to convert a number to a string in JavaScript
- a = String(n)
- a = n.toString()
- a = ““+n, so combining num with string is easy
new_str = 'hello world' + num
SVG drawing
Research a lot for SVG draw with javascript.
- How to draw an arc in SVG like
.arcTo()in canvas?- use
to draw arc, such as ` ` M = moveto // necessary L = lineto // necessary Q = quadratic Belzier curve // not used A = elliptical Arc // good! - Spent much time and learned, A is better than Q, and
Ais very similar withcanvas.arcTo():d="M x1 y1 A rx ry, x-axis-rotation, large-arc-flag,sweep-flag, x2 y2" // arc is a part of an eclipse with rx,ry and rotated, starts from (x1,y1) and ends at (x2,y2), small arc if large-arc-flag== 0, colokwise arc if sweep-flag == 1 canvas.moveTo(xa,ya); canvas.arcTo(cx,cy,xb,yb,r); // arc is tangent with (xa,ya)-(cx,cy) and (cx,cy)-(xb,yb) and radius r, may not start from (xa,ya) and may not end at (xb,yb).
- use
- How to manipulate data for a
<path>element from script? Solution Source- use
path.setAttribute()or useSVG DOM - set the whole path with
path.setAttribute('d','M150,0 L150,100 200,300 Z') - set partial value like
path.pathSegList.getItem(2).y = -10 - generate a path with
SVGPathElementvar _l = 10; var pathEl = document.createElementNS("http://www.w3.org/2000/svg", "path"); pathEl.setAttribute('d','M'+_l+' 100 Q 100 300 '+_l+' 500' ); ISVGPathSegArcAbs retVal = object.createSVGPathSegArcAbs(in float x, in float y, in float r1, in float r2, in float angle, in boolean largeArcFlag, in boolean sweepFlag)
- use
- place your
<script>elements at the bottom of yoursvgdocument. Alternatively, you can create a callback function at the top of your document that is only invoked when the rest of the document is ready:
Back to subcontents RNA web demo
12/30/2017 Sat
New Request:
- (☆☆☆☆☆) In P/R/F-beam plot, remove the F-score series. It became a P/R-beam plot.
- (★★★★☆) In P/R-beam plot, add a switch to let the user choose between log-scale and linear-scale. (default: log)
- Time consuming, but it feels good when I finish it. The position of button is annoying. Finally I remove the designated size of the button.
- Since x is number in log view, the
dataTableis easy, while x is string in linear view, so I need another dataTable ready for log/linear view switch.
- (★★★☆☆) In P/R-beam plot, show both PPV and sensitivity instead of just one with mouse hover. ‘This might be difficult.’
- Learnt a concept named
tooltips, displaying point values when a mouse hover on an element/button/plot point. - In google charts lib, tooltips are defaulted as
x_value \n y_series_label: y_value. Now I want to show all series in 1 x value and show the x label.

- Find an example on google developers after a really long search, in the section of
multi-domain chart. But true solution is found in the example ofLondon Olympics Medalsframe from another google chart guide-Tooltips, along with the definition of Options Configuration:var options = { ... colors: ['#FFD700', '#C0C0C0', '#8C7853'], // assign colors for each series, or do it in 'series' focusTarget: 'category', // This line makes the entire category's tooltip active. 'category' - Focus on a grouping of all data points along the major axis. Correlates to a row in the data table. ... }; - Here is the result I’m satisfied with:
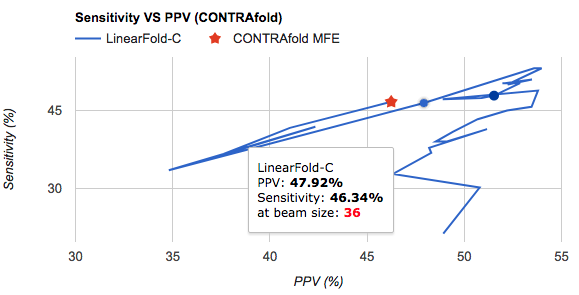
- Learnt a concept named
- (★★★★★) In P/R-beam plot, use “b=1” instead of “1” in mouse hover(tooltip).
- I can use the
categoryoption to list all series on hover, but x-label(domain label) is still missing from default.
- I can use the
- (★☆☆☆☆) In P/R-beam plot, add a PPV line and a Sensitivity line for CONTRAfold/Vienna.
- Done before requirement received in email.
- (☆☆☆☆☆) In R-P plot, connect the points with a line, change
.ScatterChart()toLineChart(). - (★★★★☆) In R-P plot, show beam size, besides the sens and ppv, in mouse hover.
- Edit a lot for the tooltip(mouse hover) in R-P plot. If I want to add
beam size, which is not shown in the graph, displayed in tooltip on each point, I need to creat a customized HTML as tooltip string. - Not only to add
'p': {'html': true}in creating.addColumn({'type': 'string', 'role': 'tooltip', 'p': {'html': true}}), to addtooltip: { isHtml: true }inoptions, but also remember to addfocusTarget: 'category'inoptions, this line makes the entire category’s tooltip active, for both HTML-based customized tooltip and default tooltip display in multi-series.

- Scripting an HTML is annoying.
- Edit a lot for the tooltip(mouse hover) in R-P plot. If I want to add
- (★★★★★)(4 hours) In both plots, highlight the corresponding points when the user slides the sliding bar.
- Find a solution in the section of
Customizing individual pointsin google chart guide-Customizing Points. - Read carefully and try millions times for the individual point display.
- When it is
.Linechart(), points are not shown by default, even there is an assigned customized individual point. Only adding sth likepointSize: 7,inoptionscan make points displayed. In fact, all the points will be that size, which is ulgy in alinechart, I have to minimize the pointSize to resume a clear line chart. However, this size is also the one for hover-highlighted point size. So there is a trade-off. Sigh…(This problem doesn’t appear one day later even no pointSize given. ???) - Customizing individual point in P/R-beam plot goes well. Spent some time in choosing point style(star/sides/dent, ect.). However, I spent 3 hours in debugging individual point customization in R-P plot. Why?
- In P/R-beam plot, beam starts from 1 and rowIndex starts from 0. Everything is OK.
dataTable.setValue(rowIndex-1, 2, style); // [beam, P, null, R, null, P_fix, R_fix] --> [beam, P, style, R, null, P_fix, R_fix] dataTable.setValue(rowIndex-1, 4, style); // [beam, P, style, R, null, P_fix, R_fix] --> [beam, P, style, R, style, P_fix, R_fix] - In R-P plot, beam starts from 1 and rowIndex starts from 1, there is a fixed point (CONTRAfold MFE) at the top of row, so beam data starts from 1.
dataTable.setValue(rowIndex, 2, style); // [beam, P, null, R, null, P_fix, R_fix] --> [beam, P, style, R, null, P_fix, R_fix] dataTable.setValue(rowIndex, 4, style); // [beam, P, style, R, null, P_fix, R_fix] --> [beam, P, style, R, style, P_fix, R_fix]There was an error running this code. I spent a lot of time to figure it out:
rowIndexis a string instead of a number, which comes fromdocument.getElementById("beamslidebar").value(what??? beamslidebar is a slide bar using standard ‘range’).rowIndexwill remain a string, not be converted to a number, until operated with a math (like rowIndex+=1). New find!
- In P/R-beam plot, beam starts from 1 and rowIndex starts from 0. Everything is OK.
- Find a solution in the section of
Back to subcontents RNA web demo
12/31/2017 Sun
My own update
- Adjust the position of plots to left, solving the problem of overlaping, by transparent ground with
backgroundColor: { fill:'transparent' }inoptions. Re-write the functionplot_411, combine two dataTable instead of generating two seperately.- If I want to display pure linear x axis(beam size) from 1-800, I can use the same dataTable for both log and linear view.
- If I want to enlarge the range of major range (1-200) and shrink (300-800), I can map 300->201,400->201,..,800->206, and make (1-206) linear. Then use
ticks:[{v:201,f:'300'},...]in hAxis iption to make x label right.
- Regroup functions to a bettle structure.
- Solve the unchecked item in log, ‘(★★★★★) In P/R-beam plot, use “b=1” instead of “1” in mouse hover(tooltip).’
- In order to list all series on hover, as well as x-label(domain label), I need to abandon the default
categorydisplay. The only way I figured out is customized HTML as tooltip string, the same as used in R-P plot.data_C_1.addRow([beam, createCustomHTMLContent_1(beam, data_C_1.getColumnLabel(2), data_C_1.getColumnLabel(4), data_C_1.getColumnLabel(6), data_C_1.getColumnLabel(7), P, R, P_fix, R_fix), P, , R, , P_fix, R_fix]);The second element in a row is a tooltip, added as a column header
data_C_1.addColumn({'type': 'string', 'role': 'tooltip', 'p': {'html': true}}); - In the function
createCustomHTMLContent_1, I editted a new html line, including series line color, with times of trails of hex colors. - Here is the result I’m satisfied with:
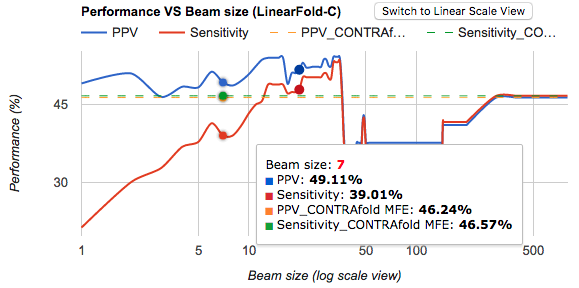
- In order to list all series on hover, as well as x-label(domain label), I need to abandon the default
Back to subcontents RNA web demo
01/01/2018 Mon
Problem shooting
My call to google.visualization.DataTable() returns “Uncaught TypeError: google.visualization.DataTable is not a constructor(…)”.
On the first click I got the error and no chart, but on the second click the page would display the chart correctly without any errors.
The same problem discussed on Google Groups.
My function was `
google.charts.setOnLoadCallback(load_draw_go(d,R,circleScale,halfOpen));`
It should be:
google.charts.setOnLoadCallback(function(){load_draw_go(d,R,circleScale,halfOpen)}); // draw graphs and plots on the right
function load_draw_go(d,R,circleScale,halfOpen){} // define the function
...
By naming your callback function and giving it parameters, you are actually calling it at that time and only the result of that function call will be passed in to the setOnLoadCallback. If you call your draw function before the library is loaded, then many things will be undefined, such as the DataTable constructor. By just naming the callback function (e.g. just drawChart), you are passing in a reference to that function. So if you want to pass parameters to your drawChart function, you need to wrap that call in another function declaration, like this:
google.charts.load(‘current’, {packages: [‘corechart’, ‘line’]);
google.charts.setOnLoadCallback( function() { drawChart(loc) });
function drawChart(loc) { … }
Back to subcontents RNA web demo
01/02/2018 Tue
- Meeting with Prof. Huang and Dezhong, discussed a little about the web demo, then asked me to read about R&E (
$n^6$) and D&P ($n^5$), then run the corresponding software ? and NUPACK. - Prof. Huang felt the webpage may freeze when we slide to much. I explained with console log that each move on slide bar, beam value is updated then data file loaded and graphs/plots are rendered.
- At home, I tried two methods to speed up the demo for frequent silde.
- I tried to load data at the beginning, save it to a
globalvariable (pairingList). Then the draw module and plot module can use the current pairingList to finish the job. Codes are updated and rearranged. But I found that the beam value is always the same as last click, not from current click. - After variable monitoring in console log, I got a conclusion, which seemed to be in my sight days ago, that
$.getJSON(url)and$.get(url)are executed after the whole page is loaded. Here is the example. Line 59 is the first run in console, with previous seq length referred by beam value, then line 38, and finally line 52, with the correct seq length referred by the current beam value.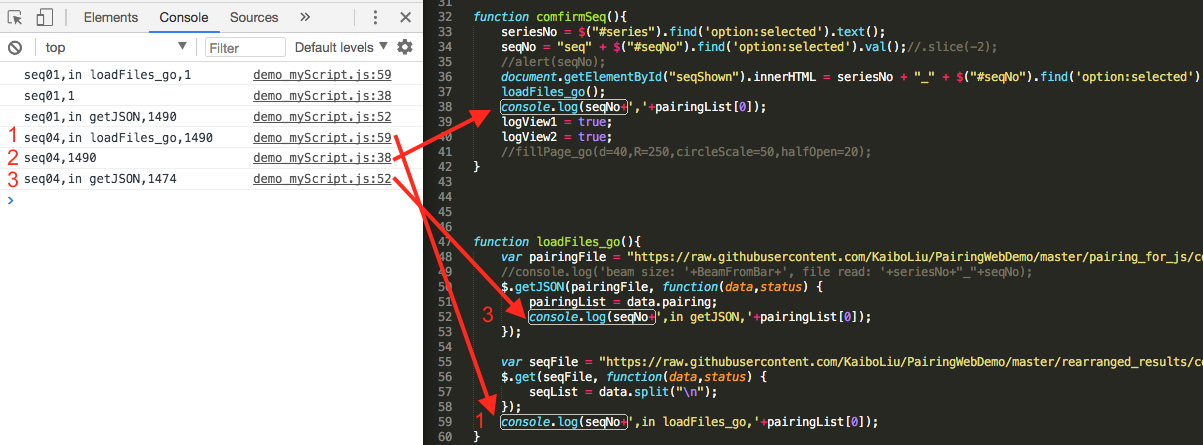
- So I cannot use
$.getJSON(url)or$.get(url)to save data globally. I can only manipulate the data from file within the function upon$.getJSON(url). - The last option I have, to avoid lagging refresh speed on webpage, is to change the attr event of slide bar(
rangetype ininputtag of html5) fromoninput=function()toonchange=function(). The former is triggered simultaneously when we slide the bar (value inputted but mouse not released), while the latter is triggered when we finish sliding the bar (mouse click released).
- I tried to load data at the beginning, save it to a
- (★★★★☆)(3 hours) Spent much time to enable re-draw when clicking on a chart/plot point with certain beam size. Digging in Google guide-event. The solution is
addListeneronselectevent.// The select handler. Call the chart's getSelection() method function selectHandler() { var selectedItem = thisChart.getSelection()[0]; if (selectedItem) { //var value = data.getValue(selectedItem.row, selectedItem.column); BeamFromBar = selectedItem.row + 1; // 1-206, need to be converted to 1-200,300,400,500,600,700,800 document.getElementById("beamslidebar").value = BeamFromBar; // update the position on beam size slide bar if (BeamFromBar > 200) BeamFromBar = (BeamFromBar - 200) * 100 + 200; // do other things, like re-draw } } // Listen for the 'select' event, and call my function selectHandler() when the user selects something on the chart. google.visualization.events.addListener(thisChart, 'select', selectHandler);
Back to subcontents RNA web demo
Back to subcontents Pseudoknot
01/04/2018 Thu
- Have trouble getting files from server end (forbidden:403), resume to get files from github end.
Back to subcontents RNA web demo
- Add another family(series) named
grp1.01/09/2018 Tue
01/17/2018 Wed
- learn from encorechow’s innotator project
- check ip for
flopandironcreekwithip addr show. I can only pingironcreekwithinflop, not from home directly.
| server | ip |
|---|---|
flop.engr.oregonstate.edu |
128.193.54.168/23(ip addr show)/ 73.67.241.185(log in, ping able) |
flip.engr.oregonstate.edu |
128.193.36.41/22(ip addr show and ping) |
ironcreek.eecs.oregonstate.edu |
128.193.38.74/22(ping able in flop) |
web.engr.oregonstate.edu |
128.193.40.12(ping able) |
shell.onid.oregonstate.edu |
128.193.4.170(not ping able) |
- learn flask for web develop(request and response)
- install flask in virtual environment
virtualenv --system-site-packages ~/.virtualenvs/flask workon flask pip install flask - version of flask
0.12.2 - logger in flask
- auto-submit an upload form when a file is selected (skip clicking submit button)
$('#file').change(function() { $('#target').submit(); });or, in the form with onchange event :
<form action="upload.php" method="post" enctype="multipart/form-data"> <input type="file" name="filename" onchange="form.submit()"> </form> enctype="multipart/form-data"in form tag is very important- However, the submission method to flask will make the whole page refresh. So I need ajax
- install flask in virtual environment
Back to subcontents RNA web demo
01/18/2018 Thu
- explore communication between
flipandironcreek - code location is
/scratch/dengde/fastdecode/fastcky_rerun/build_beamckypar, we need to run a script~dengde/run_beamckypar_new.shto use specific version of gcc. - line break in the attr
placeholderin: use ` ` instead of `\n` - terminologies in flask.(source)
- PythonDecorato
- view function: return string, no int or float
- Variable in
<>of@app.route()should be te same as the parameter view function url_for()pass the name of a view function as string parameter, return the url of the corresponding function.- good material: flask设计思路
Back to subcontents RNA web demo
01/19/2018 Fri
- What’s the difference between
python hello.pyandflask run```bash $ export FLASK_APP=hello.py $ flask run- Running on http://127.0.0.1:5000/ ``` Back to subcontents RNA web demo
01/21/2018 Sun
- Check if the uploaded file is allowed:
- html form: ```
```python ALLOWED_EXTENSIONS = set(['txt', 'pdf', 'png', 'jpg', 'jpeg', 'gif']) def allowed_file(filename): return '.' in filename and filename.rsplit('.', 1)[1] in ALLOWED_EXTENSIONS
Back to subcontents RNA web demo
01/29/2018 Mon
- Finish data transmission between flip and ironcreek with
socketin python.
graph LR
V(<b>browser</b><br>web.engr.ore.../onid<br>128.193.40.12/onid)-->|visit <b>but</b> <br>no data upload|P(<b>./public_html/</b><br>html,css,js<br>)
P-->|in|D(<b>flop</b><br><b>ip</b>: 73.67.241.185)
P-->|in|B(<b>flip/access</b><br><b>host</b>: flip3.engr.oreg...<br><b>ip</b>: 128.193.36.41)
P-->|in|A(<b>ironcreek</b><br><b>host</b>:ironcreek.eecs...<br><b>ip</b>: 128.193.38.74<br><br><b>do the math</b>)
V-->|ssh|D
subgraph server data flow
A
B
D
P
end
subgraph client off campus
V
end
classDef green fill:#9f6,stroke:#333,stroke-width:2px;;
classDef orange fill:#f48c42,stroke:#f66,stroke-width:2px,stroke-dasharray: 5, 5;
class A,V,B,D green
%%class P orange
01/30/2018 Wed
read files with jQuery in local html, there is an error: Cross origin requests are only supported for protocol schemes: http, data, chrome, chrome-extension, https. I found a solution onlie:
Just to be explicit - Yes, the error is saying you cannot point your browser directly at file://some/path/some.html
- Python 2
If you have Python installed… Change directory into the folder where your file some.html or file(s) exist using the command cd /path/to/your/folder Start up a Python web server using the command python -m SimpleHTTPServer
This will start a web server that hosts your entire directory listing, which will be made accessible through the following URL: http://localhost:8000
This approach is built in to any Python installation.
- Python 3
Do the same steps, but use the following command instead python3 -m http.server
Back to subcontents RNA web demo
02/02/2018 Fri
sequenceDiagram
participant C as client off campus
participant S1 as flip/access
participant S2 as ironcreek
%note left of C: visit
note over C: visit web.engr.../~onid
C->>S1:customized seq
alt file upload
C->>S1: file request
else name & seq input
C->>S1: text request
end
note over S1: write seq to file save SeqName
S1->>S2: call prog with file name
note over S2: run linear C/V write to 'pairing'
S2->>S1:return the message 'done'
note over S1: render a new page with 'pairing'
S1->>C: redirect to the new page
note over C: Wow!